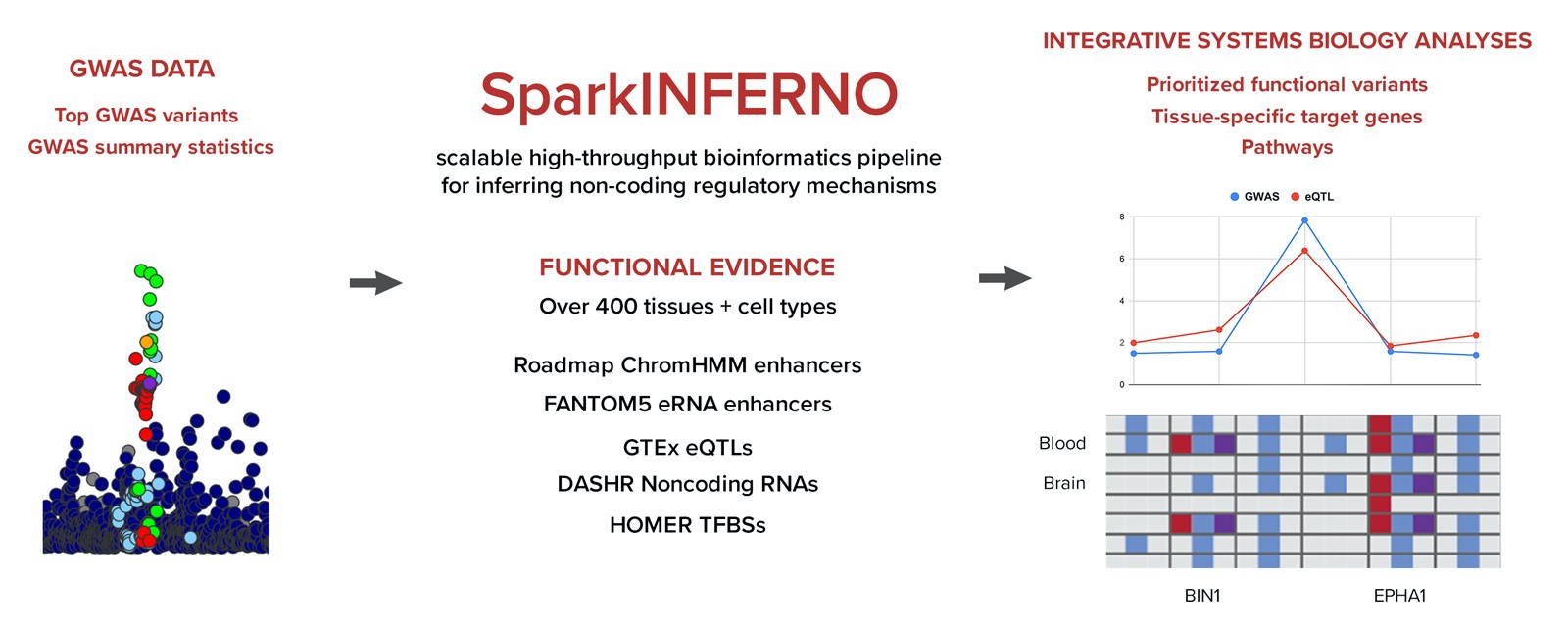
PNGC is pleased to announce the release of SparkINFERNO, a next-generation of INFERNO (INFERring the molecular mechanisms of NOn-coding genetic variants) bioinformatics pipeline.
While genome-wide association studies (GWASs) have identified over 70,000 disease-associated genetic variants, most hits are in the non-coding region, which makes their interpretation difficult. SparkINFERNO, a scalable, high-throughput bioinformatics pipeline characterizing noncoding GWAS findings, helps to address this issue. The SparkINFERNO algorithm integrates GWAS summary statistics with large-scale collection of functional genomics datasets spanning enhancer activity, transcription factor binding, expression quantitative trait loci, and other functional datasets across more than 400 tissues and cell types.
Some of the SparkINFERNO highlights include:
1) Scalable, extensible parallel analysis architecture based on Apache Spark
2) Integrated large-scale functional genomics data repository with scalable querying interface (Apache Spark + Giggle-based genomic indexing)
3) Integrated array of parallelized integrative systems biology analyses to identify tissue contexts, target genes and downstream biological processes underlying GWAS findings
SparkINFERNO source code is available here: https://bitbucket.org/wanglab-upenn/SparkINFERNO. Pre-built SparkINFERNO Docker image with integrated Apache Spark environment is available here: https://hub.docker.com/r/wanglab/spark-inferno.
SparkINFERNO has been published in Bioinformatics. The full manuscript can be found here: https://doi.org/10.1093/bioinformatics/btaa246.
PNGC congratulates authors Pavel P. Kuksa, Chien-Yueh Lee, Alexandre Amlie-Wolf, Prabhakaran Gangadharan, Elizabeth Mlynarski, Yi-Fan Chou, Han-Jen Lin, Heather Issen, Emily Greenfest-Allen, Otto Valladares, Yuk Yee Leung, and Li-San Wang on this accomplishment!